|
cloudy
trunk
|
|
cloudy
trunk
|
#include "cdstd.h"#include "cddefines.h"#include "prt.h"#include "version.h"#include "hmi.h"#include "conv.h"#include "thermal.h"#include "grainvar.h"#include "ionbal.h"#include "opacity.h"#include "radius.h"#include "atmdat.h"#include "trace.h"#include "deuterium.h"#include "grains.h"#include "mole_priv.h"#include "gammas.h"#include "mole.h"#include "freebound.h"#include "dense.h"#include "phycon.h"#include "doppvel.h"#include "rfield.h"#include "secondaries.h"#include "hextra.h"#include "rt_escprob.h"#include "iso.h"#include "h2.h"
Go to the source code of this file.
Macros | |
| #define | FLTEQ(a, b) (fabs((a)-(b)) <= 1e-6*fabs((a)+(b))) |
Enumerations | |
| enum | { UDFA = 0 } |
| enum | |
| enum | { BUFSIZE =256 } |
Variables | |
| static realnum | albedo = 0.5 |
| static map< formula_species, count_ptr< udfa_reaction > > | udfatab |
| #define FLTEQ | ( | a, | |
| b | |||
| ) | (fabs((a)-(b)) <= 1e-6*fabs((a)+(b))) |
Definition at line 2851 of file mole_reactions.cpp.
Referenced by compare_udfa(), and parse_udfa().
| anonymous enum |
| Enumerator | |
|---|---|
| UDFA | |
Definition at line 46 of file mole_reactions.cpp.
| anonymous enum |
| Enumerator | |
|---|---|
| BUFSIZE | |
Definition at line 2790 of file mole_reactions.cpp.
| anonymous enum |
Definition at line 1715 of file mole_reactions.cpp.
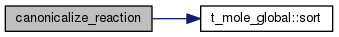
| STATIC void canonicalize_reaction | ( | count_ptr< mole_reaction > & | rate | ) |
Definition at line 2535 of file mole_reactions.cpp.
References DEBUG_ENTRY, mole_reaction::label, molecule::label, mole_reaction::nproducts, mole_reaction::nreactants, mole_reaction::products, mole_reaction::reactants, and t_mole_global::sort().
Referenced by canonicalize_reaction_label(), and newreact().
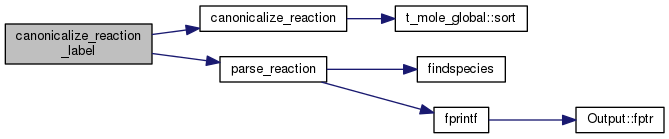
| STATIC string canonicalize_reaction_label | ( | const char | label[] | ) |
Definition at line 2519 of file mole_reactions.cpp.
References canonicalize_reaction(), DEBUG_ENTRY, and parse_reaction().
Referenced by mole_create_react(), mole_findrate_s(), and mole_generate_isotopologue_reactions().
| STATIC void compare_udfa | ( | const count_ptr< mole_reaction > & | rate | ) |
Definition at line 2953 of file mole_reactions.cpp.
References mole_reaction::a, mole_reaction::b, mole_reaction::c, CONFLICT, CORRECT, FLTEQ, MAXPRODUCTS, MAXREACTANTS, mole_reaction::products, mole_reaction::reactants, and mole_reaction::udfastate.
Referenced by newreact().
| double frac_H2star_hminus | ( | void | ) |
Definition at line 699 of file mole_reactions.cpp.
References h2, t_hmi::H2_forms_hminus, t_hmi::H2star_forms_hminus, hmi, diatomics::lgEnabled, diatomics::lgEvaluated, t_hmi::lgH2_Chemistry_BigH2, and SDIV().
Referenced by mole_h_rate_diagnostics().

| STATIC char * getcsvfield | ( | char ** | s, |
| char | c | ||
| ) |
Definition at line 2941 of file mole_reactions.cpp.
Referenced by parse_base(), and parse_udfa().
| double hmirat | ( | double | te | ) |
hmirat computes radiative association rate for H-
| te |
Definition at line 1632 of file mole_reactions.cpp.
References DEBUG_ENTRY, phycon, t_phycon::sqrte, t_phycon::te001, t_phycon::te003, t_phycon::te01, t_phycon::te03, t_phycon::te10, t_phycon::te20, and t_phycon::te70.
Referenced by mole_h_reactions().
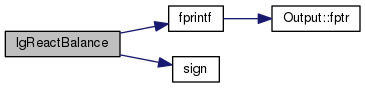
| STATIC bool lgReactBalance | ( | const count_ptr< mole_reaction > & | rate | ) |
Definition at line 2745 of file mole_reactions.cpp.
References molecule::charge, fprintf(), mole_reaction::label, molecule::nNuclide, mole_reaction::nproducts, mole_reaction::nreactants, mole_reaction::products, mole_reaction::reactants, and sign().
Referenced by newreact().
| STATIC bool lgReactionTrivial | ( | count_ptr< mole_reaction > & | rate | ) |
Definition at line 2568 of file mole_reactions.cpp.
References DEBUG_ENTRY, mole_reaction::nproducts, mole_reaction::nreactants, mole_reaction::products, and mole_reaction::reactants.
Referenced by newreact().
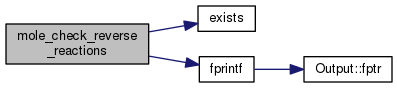
| STATIC void mole_check_reverse_reactions | ( | void | ) |
Definition at line 2201 of file mole_reactions.cpp.
References DEBUG_ENTRY, exists(), fixit, fprintf(), ioQQQ, t_trace::lgTraceMole, mole_priv::reactab, and trace.
Referenced by mole_create_react().
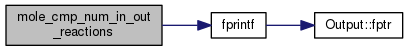
| void mole_cmp_num_in_out_reactions | ( | void | ) |
Definition at line 2307 of file mole_reactions.cpp.
References CHARS_SPECIES, DEBUG_ENTRY, fprintf(), molecule::index, ioQQQ, t_mole_global::list, mole_global, mole_reaction::nproducts, mole_reaction::nreactants, t_mole_global::num_total, mole_reaction::products, mole_priv::reactab, and mole_reaction::reactants.
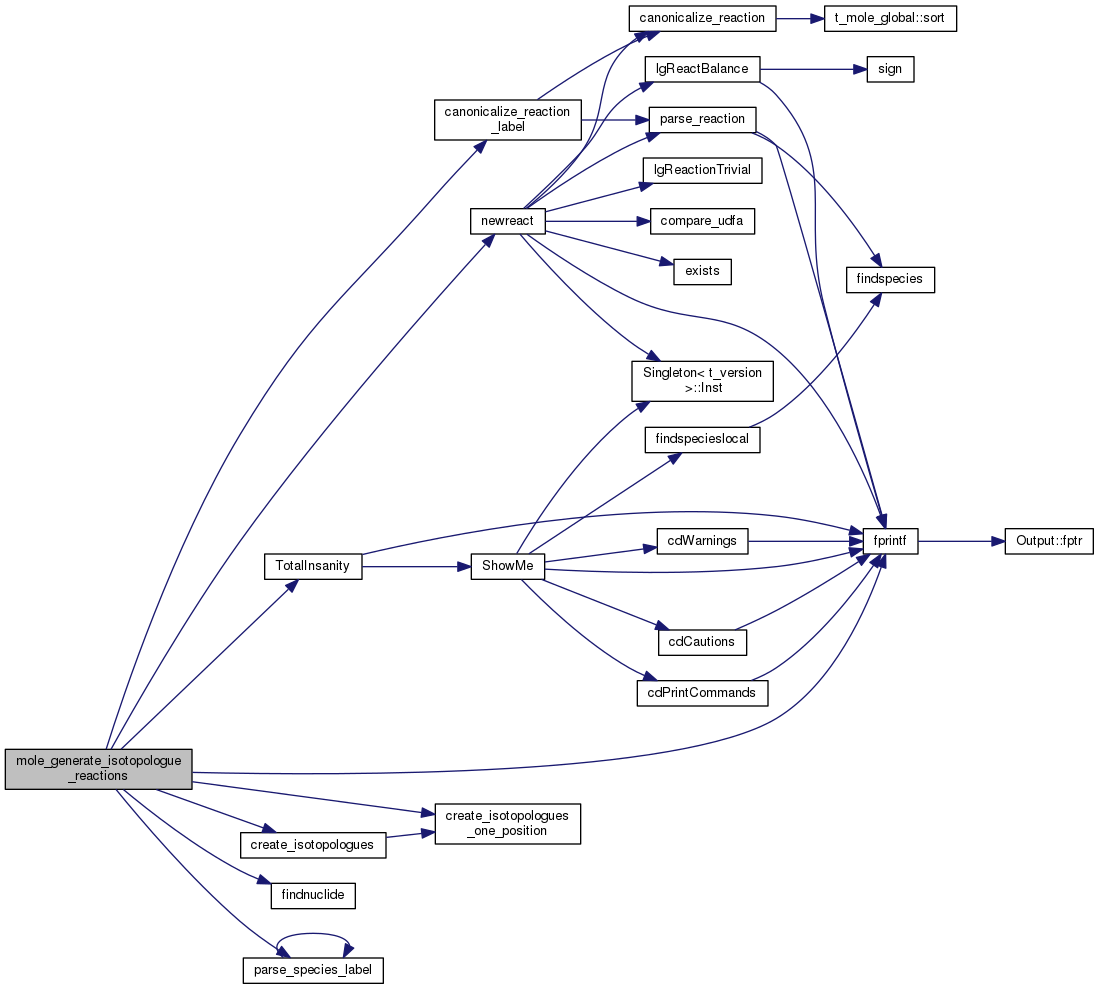
| void mole_create_react | ( | void | ) |
mole_create_react build reaction structures
Definition at line 1722 of file mole_reactions.cpp.
References canonicalize_reaction_label(), cdEXIT, DEBUG_ENTRY, dense, deut, EXIT_FAILURE, fixit, fprintf(), hmi, ipHELIUM, t_deuterium::lgElmtOn, t_dense::lgElmtOn, t_mole_global::lgFederman, t_hmi::lgLeiden_Keep_ipMH2s, t_mole_global::lgLeidenHack, t_mole_global::lgProtElim, mole, mole_check_reverse_reactions(), mole_generate_isotopologue_reactions(), mole_global, newreact(), nuclide_list, t_mole_global::offReactions, t_mole_local::old_zone, parse_base(), parse_udfa(), plot_sparsity(), pow(), mole_priv::reactab, t_mole_local::reaction_rks, read_data(), register_reaction_vectors(), and UDFA.
Referenced by InitSimPostparse().
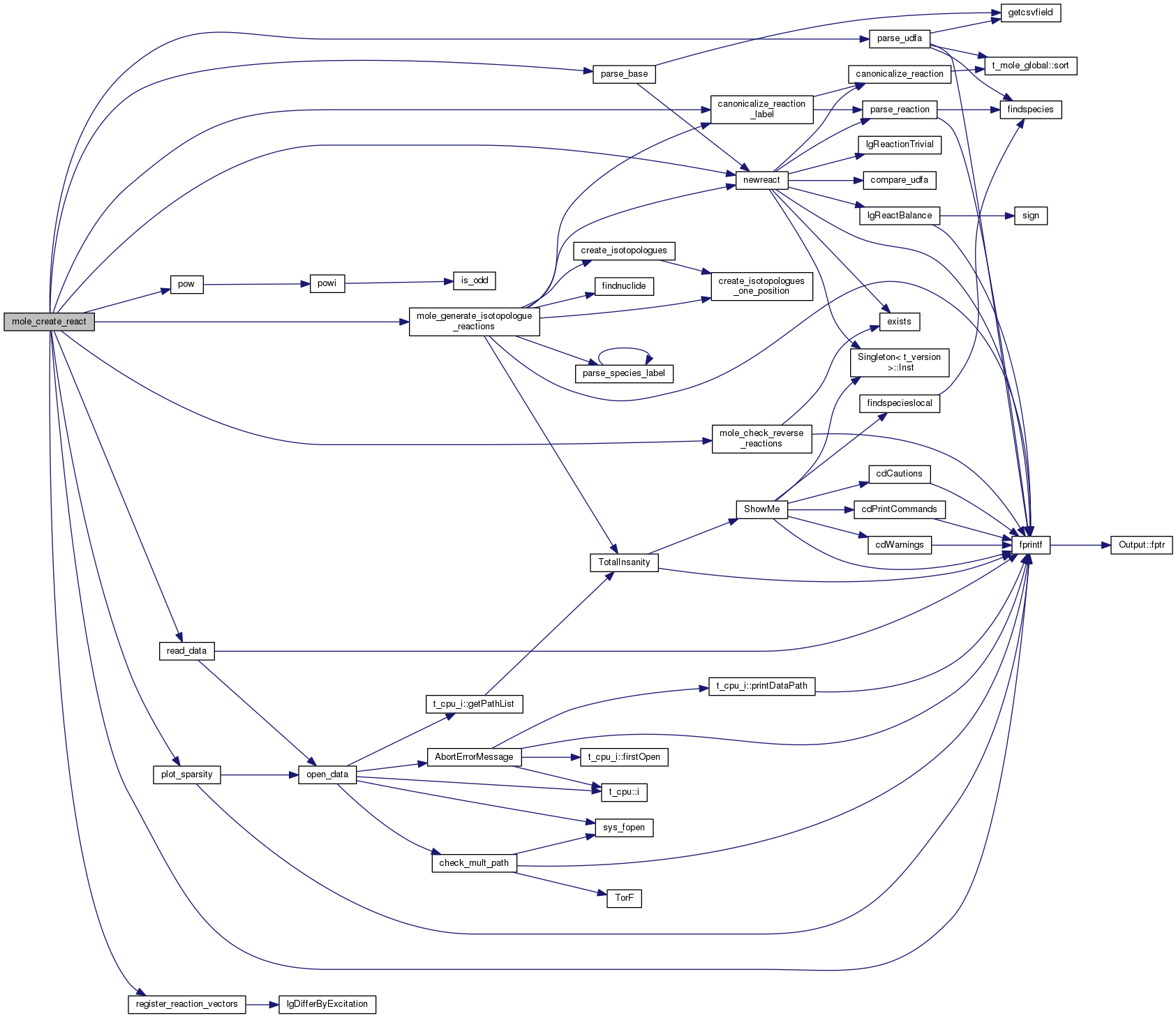
| mole_reaction* mole_findrate_s | ( | const char | buf[] | ) |
Definition at line 4123 of file mole_reactions.cpp.
References canonicalize_reaction_label(), DEBUG_ENTRY, and mole_priv::reactab.
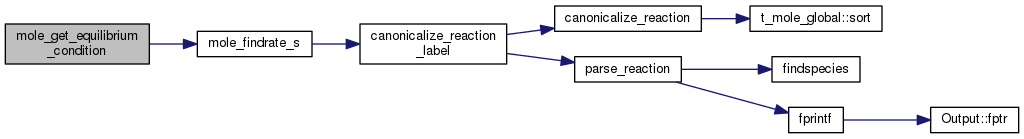
Referenced by t_mole_local::findrate(), t_mole_local::findrk(), mole_get_equilibrium_condition(), and t_mole_local::sink_rate().
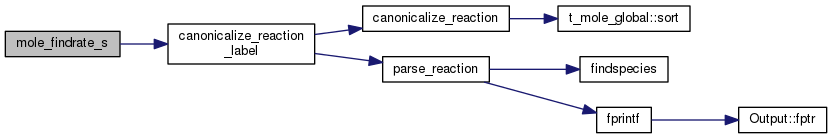
| STATIC void mole_generate_isotopologue_reactions | ( | string | atom_old, |
| string | atom_new | ||
| ) |
Definition at line 2023 of file mole_reactions.cpp.
References ASSERT, canonicalize_reaction_label(), create_isotopologues(), create_isotopologues_one_position(), DEBUG_ENTRY, findnuclide(), fixit, fprintf(), ioQQQ, newreact(), parse_species_label(), mole_priv::reactab, and TotalInsanity().
Referenced by mole_create_react().
| STATIC double mole_get_equilibrium_condition | ( | const char | buf[] | ) |
Definition at line 2243 of file mole_reactions.cpp.
References DEBUG_ENTRY, and mole_findrate_s().
| STATIC double mole_get_equilibrium_condition | ( | const mole_reaction *const | rate | ) |
Definition at line 2252 of file mole_reactions.cpp.
References BIGFLOAT, DEBUG_ENTRY, MIN2, mole_partition_function(), mole_reaction::nproducts, mole_reaction::nreactants, mole_reaction::products, mole_reaction::reactants, and SEXP_LIMIT.

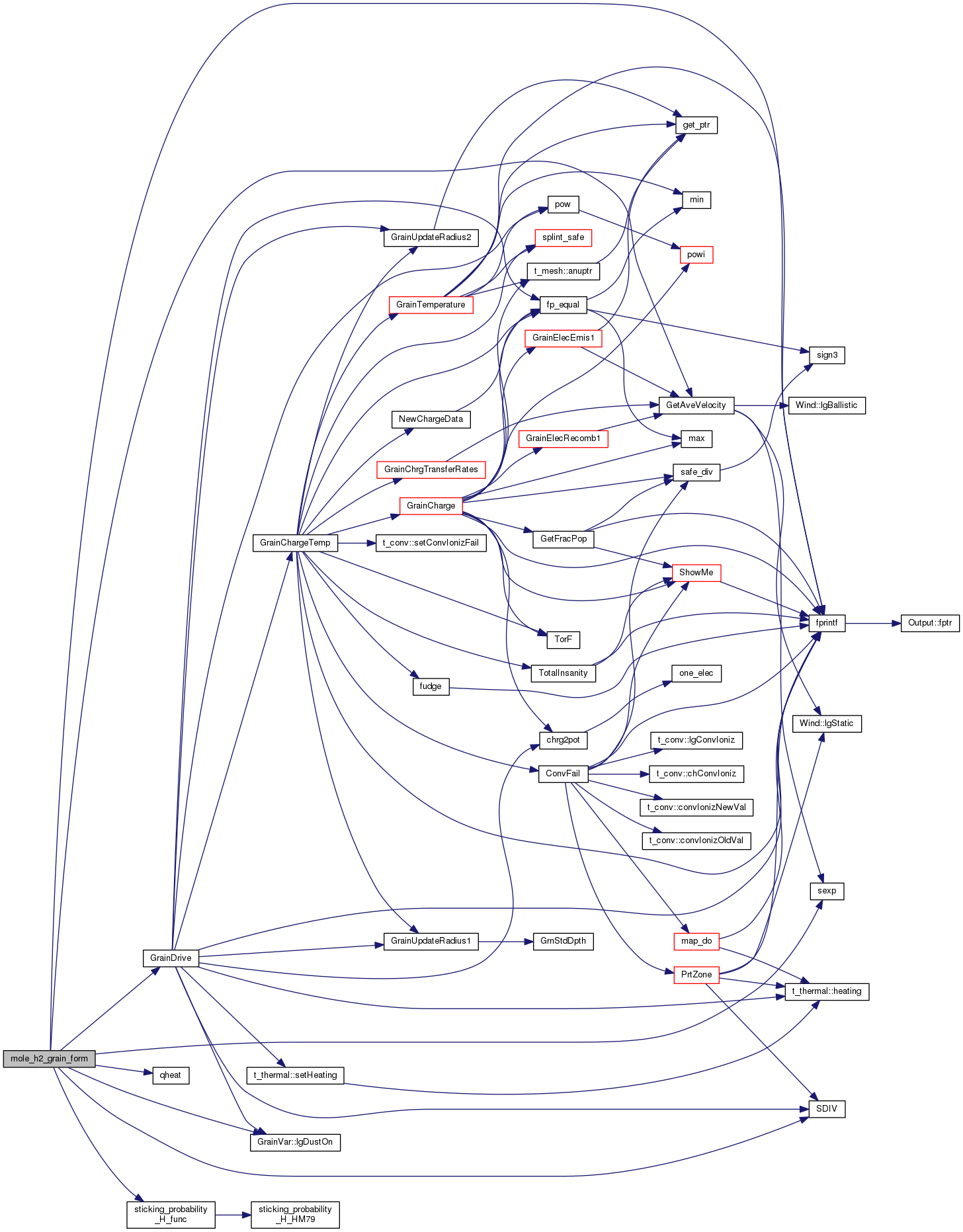
| STATIC void mole_h2_grain_form | ( | void | ) |
Definition at line 3081 of file mole_reactions.cpp.
References ASSERT, t_dense::AtomicWeight, GrainVar::bin, t_hmi::chJura, DEBUG_ENTRY, dense, ENABLE_QUANTUM_HEATING, fixit, fnzone, fprintf(), t_dense::gas_phase, GetAveVelocity(), GrainDrive(), gv, h2, hmi, ioQQQ, ipHYDROGEN, GrainVar::lgDustOn(), GrainVar::lgGrainPhysicsOn, diatomics::lgH2_grain_deexcitation, MAT_CAR, MAT_CAR2, MAT_PAH, MAT_PAH2, MAT_SIC, MAT_SIL, MAT_SIL2, NQGRID, phycon, POW2, qheat(), diatomics::rate_grain_J1_to_J0, diatomics::rate_grain_op_conserve, t_hmi::rate_h2_form_grains_set, GrainVar::rate_h2_form_grains_used_total, t_hmi::ScaleJura, SDIV(), sexp(), t_phycon::sqrte, sticking_probability_H_func(), t_hmi::Tad, and t_phycon::te.
Referenced by mole_update_rks().
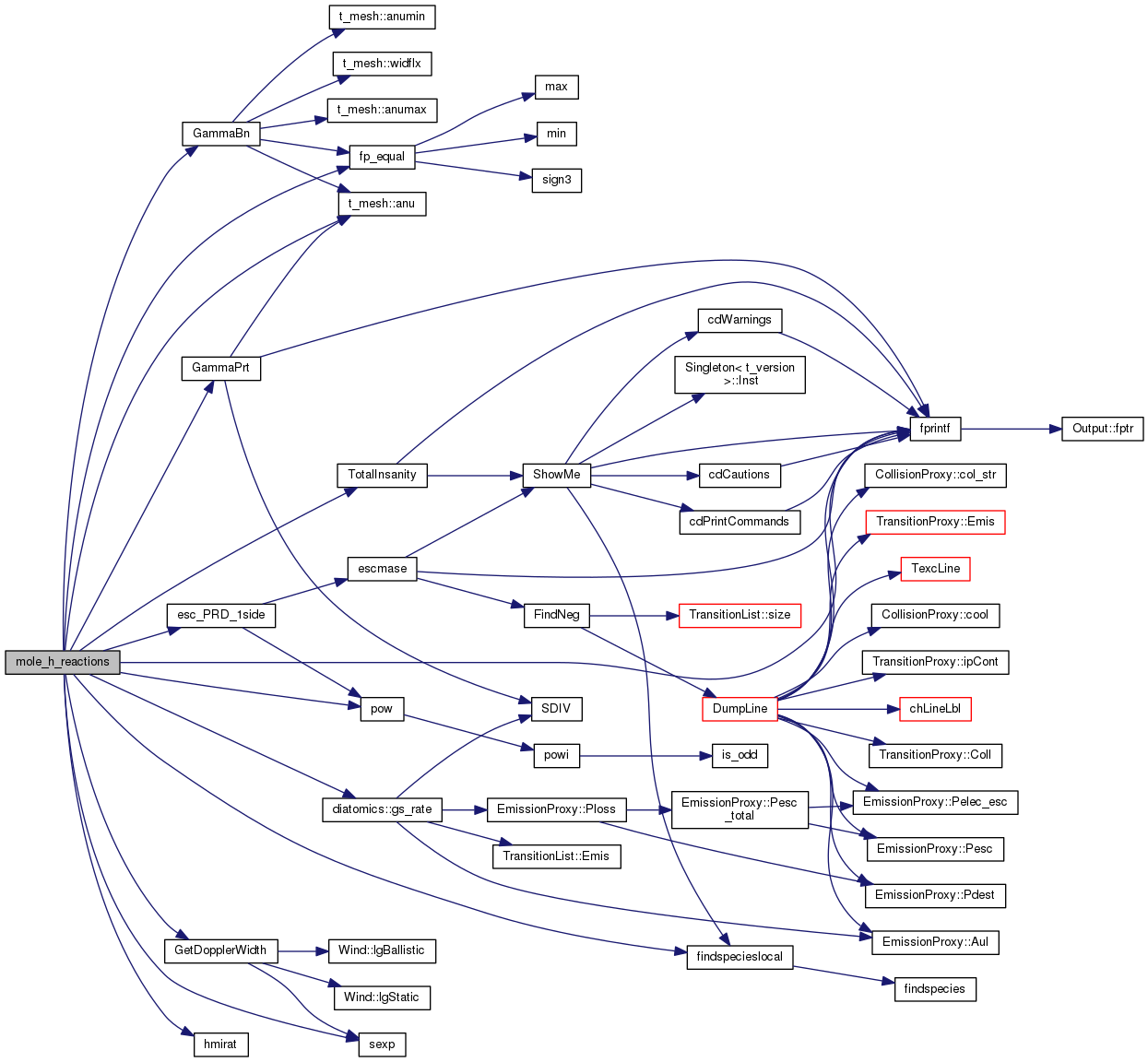
| STATIC void mole_h_reactions | ( | void | ) |
hmole_reactions - evaluates hydrogen chemistry reactions
Definition at line 3515 of file mole_reactions.cpp.
References t_mesh::anu(), ASSERT, t_dense::AtomicWeight, t_hmi::chH2_small_model_type, molezone::column, t_rfield::ConInterOut, diatomics::Cont_Dissoc_Rate_H2g, diatomics::Cont_Dissoc_Rate_H2s, conv, DEBUG_ENTRY, molezone::den, dense, diatoms, t_dense::eden, esc_PRD_1side(), t_hmi::exphmi, t_rfield::extin_mag_V_point, t_iso_sp::fb, findspecieslocal(), fixit, t_rfield::flux, fnzone, fp_equal(), fprintf(), GammaBn(), GammaPrt(), GetDopplerWidth(), diatomics::gs_rate(), h2, t_hmi::H2_H2g_to_H2s_rate_BD96, t_hmi::H2_H2g_to_H2s_rate_BHT90, t_hmi::H2_H2g_to_H2s_rate_ELWERT, t_hmi::H2_H2g_to_H2s_rate_TH85, t_hmi::H2_H2g_to_H2s_rate_used, t_hmi::H2_photodissoc_BHT90, t_hmi::H2_photodissoc_ELWERT_H2g, t_hmi::H2_photodissoc_ELWERT_H2s, t_hmi::H2_photodissoc_TH85, t_hmi::H2_photodissoc_used_H2g, t_hmi::H2_photodissoc_used_H2s, t_hmi::H2_Solomon_dissoc_rate_BD96_H2g, t_hmi::H2_Solomon_dissoc_rate_BD96_H2s, t_hmi::H2_Solomon_dissoc_rate_BHT90_H2g, t_hmi::H2_Solomon_dissoc_rate_BHT90_H2s, t_hmi::H2_Solomon_dissoc_rate_ELWERT_H2g, t_hmi::H2_Solomon_dissoc_rate_ELWERT_H2s, t_hmi::H2_Solomon_dissoc_rate_TH85_H2g, t_hmi::H2_Solomon_dissoc_rate_TH85_H2s, t_hmi::H2_Solomon_dissoc_rate_used_H2g, t_hmi::H2_Solomon_dissoc_rate_used_H2s, t_hmi::H2Opacity, hd, t_phoHeat::HeatNet, hmi, t_hmi::hmicol, t_hmi::HMinus_induc_rec_cooling, t_hmi::HMinus_induc_rec_rate, t_hmi::HMinus_photo_heat, t_hmi::HMinus_photo_rate, hmirat(), ioQQQ, t_rfield::ip1000A, t_rfield::ipG0_DB96_hi, t_rfield::ipG0_DB96_lo, t_rfield::ipG0_spec_hi, t_rfield::ipG0_spec_lo, t_rfield::ipG0_TH85_hi, t_rfield::ipG0_TH85_lo, ipHE_LIKE, ipHELIUM, t_hmi::iphmin, t_opac::iphmop, ipHYDROGEN, iso_sp, iteration, diatomics::lgEnabled, diatomics::lgEvaluated, t_hmi::lgH2_Chemistry_BigH2, t_hmi::lgLeidenCRHack, t_mole_global::lgLeidenHack, t_mole_global::lgStancil, MAX2, mole_global, t_conv::nPres2Ioniz, t_conv::nTotalIoniz, nzone, opac, t_rfield::outlin, t_rfield::outlin_noplot, diatomics::photodissoc_BigH2_H2g, diatomics::photodissoc_BigH2_H2s, phycon, pow(), POW2, t_radius::r1r0sq, radius, diatomics::rel_pop_LTE_g, t_hmi::rel_pop_LTE_H2g, t_hmi::rel_pop_LTE_H2p, t_hmi::rel_pop_LTE_H2s, t_hmi::rel_pop_LTE_H3p, t_hmi::rel_pop_LTE_Hmin, diatomics::rel_pop_LTE_s, rfield, secondaries, sexp(), SMALLFLOAT, diatomics::Solomon_dissoc_rate_g, diatomics::Solomon_dissoc_rate_s, t_opac::TauAbsFace, t_phycon::te, t_phycon::te32, TotalInsanity(), t_hmi::UV_Cont_rel2_Draine_DB96_depth, t_hmi::UV_Cont_rel2_Draine_DB96_face, t_hmi::UV_Cont_rel2_Habing_spec_depth, t_hmi::UV_Cont_rel2_Habing_TH85_depth, t_hmi::UV_Cont_rel2_Habing_TH85_face, and t_secondaries::x12tot.
Referenced by mole_update_rks().
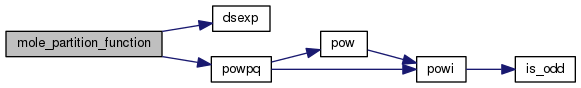
Definition at line 2282 of file mole_reactions.cpp.
References ASSERT, BIGFLOAT, DEBUG_ENTRY, dsexp(), fixit, molecule::form_enthalpy, molecule::label, molecule::mole_mass, phycon, powpq(), and t_phycon::te.
Referenced by mole_get_equilibrium_condition().
| void mole_rk_bigchange | ( | void | ) |
Definition at line 3023 of file mole_reactions.cpp.
References ASSERT, DEBUG_ENTRY, fprintf(), mole_reaction::index, ioQQQ, mole_reaction::label, mole, nzone, t_mole_local::old_reaction_rks, t_mole_local::old_zone, mole_priv::reactab, and t_mole_local::reaction_rks.
Referenced by ZoneEnd().

| void mole_update_rks | ( | void | ) |
mole_update_rks update rate coefficients, only temp part
Definition at line 2997 of file mole_reactions.cpp.
References mole_reaction::a, DEBUG_ENTRY, fprintf(), mole_reaction::index, ioQQQ, mole_reaction::label, mole, mole_h2_grain_form(), mole_h_reactions(), mole_priv::reactab, t_mole_local::reaction_rks, and mole_reaction::rk().
Referenced by mole_drive().
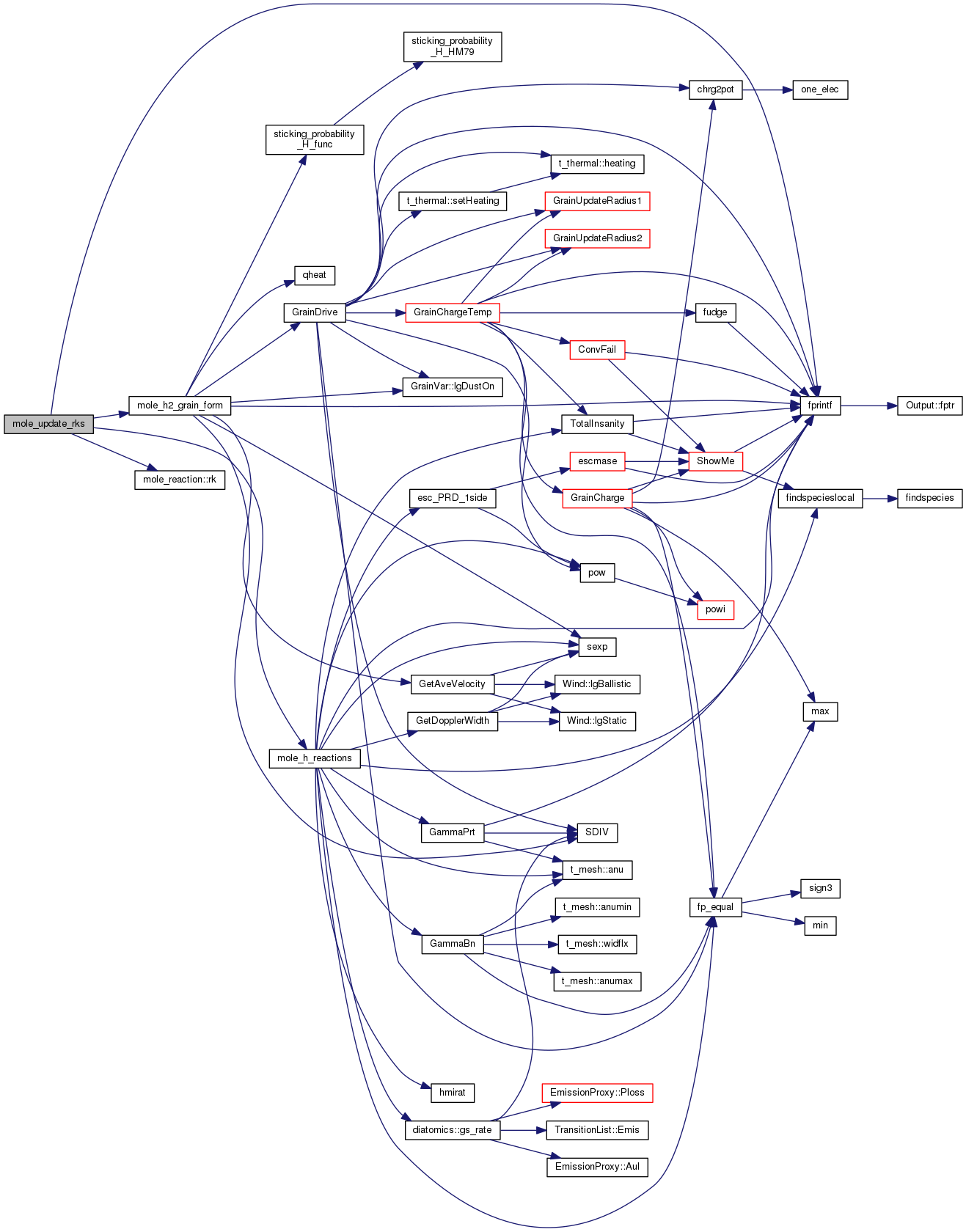
| STATIC void newreact | ( | const char | label[], |
| const char | fun[], | ||
| double | a, | ||
| double | b, | ||
| double | c | ||
| ) |
Definition at line 2361 of file mole_reactions.cpp.
References ABSENT, ASSERT, canonicalize_reaction(), cdEXIT, compare_udfa(), DEBUG_ENTRY, exists(), EXIT_FAILURE, fprintf(), Singleton< t_version >::Inst(), ioQQQ, t_phycon::lgPhysOK, t_prt::lgPrintTime, lgReactBalance(), lgReactionTrivial(), t_version::lgRelease, t_version::lgReleaseBranch, mole_global, t_mole_global::offReactions, parse_reaction(), phycon, prt, mole_priv::reactab, and UDFA.
Referenced by mole_create_react(), mole_generate_isotopologue_reactions(), and parse_base().
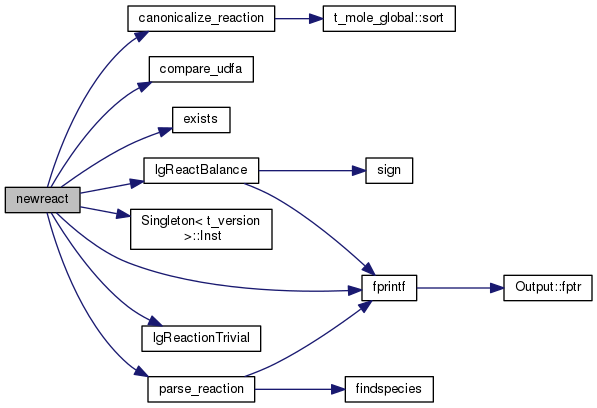
| STATIC void parse_base | ( | char * | s | ) |
Definition at line 2344 of file mole_reactions.cpp.
References getcsvfield(), and newreact().
Referenced by mole_create_react().
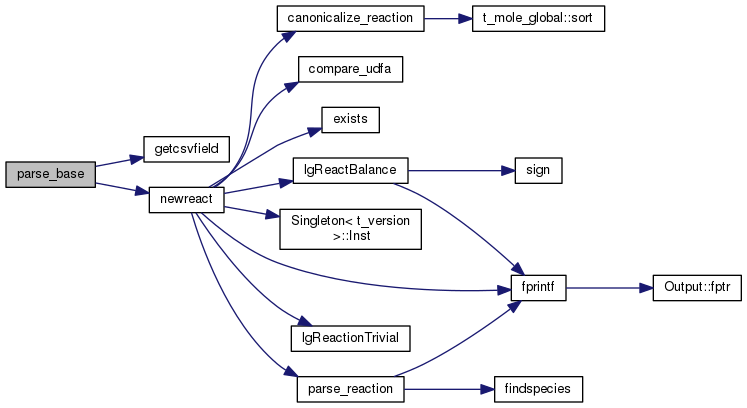
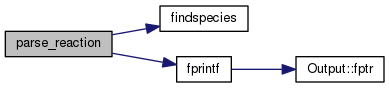
| STATIC long parse_reaction | ( | count_ptr< mole_reaction > & | rate, |
| const char | label[] | ||
| ) |
Definition at line 2439 of file mole_reactions.cpp.
References ASSERT, cdEXIT, DEBUG_ENTRY, EXIT_FAILURE, findspecies(), fprintf(), ioQQQ, molecule::isEnabled, t_trace::lgTraceMole, MAXPRODUCTS, MAXREACTANTS, molecule::n_react, mole_reaction::nproducts, mole_reaction::nreactants, null_mole, mole_reaction::products, mole_reaction::reactants, and trace.
Referenced by canonicalize_reaction_label(), and newreact().
| STATIC void parse_udfa | ( | char * | s | ) |
Definition at line 2852 of file mole_reactions.cpp.
References ASSERT, findspecies(), FLTEQ, fprintf(), getcsvfield(), MAXPRODUCTS, MAXREACTANTS, null_mole, and t_mole_global::sort().
Referenced by mole_create_react().
| STATIC void plot_sparsity | ( | void | ) |
Definition at line 2671 of file mole_reactions.cpp.
References fprintf(), molecule::index, MALLOC, mole_global, mole_reaction::nproducts, mole_reaction::nreactants, t_mole_global::num_total, open_data(), mole_reaction::products, mole_reaction::pvector, mole_priv::reactab, mole_reaction::reactants, and mole_reaction::rvector.
Referenced by mole_create_react().
| STATIC void read_data | ( | const char | file[], |
| void(*)(char *s) | parse | ||
| ) |
Definition at line 2830 of file mole_reactions.cpp.
References BUFSIZE, cdEXIT, DEBUG_ENTRY, EXIT_FAILURE, fixit, fprintf(), and open_data().
Referenced by mole_create_react().
| STATIC void register_reaction_vectors | ( | count_ptr< mole_reaction > | rate | ) |
Definition at line 2589 of file mole_reactions.cpp.
References DEBUG_ENTRY, molecule::groupnum, lgDifferByExcitation(), mole_reaction::nproducts, mole_reaction::nreactants, mole_reaction::products, mole_reaction::pvector, mole_reaction::pvector_excit, mole_reaction::reactants, mole_reaction::rvector, and mole_reaction::rvector_excit.
Referenced by mole_create_react().
| STATIC double sticking_probability_H_func | ( | double | T_gas, |
| double | T_grain | ||
| ) |
Definition at line 4453 of file mole_reactions.cpp.
References DEBUG_ENTRY, S, and sticking_probability_H_HM79().
Referenced by mole_h2_grain_form().
| STATIC double sticking_probability_H_HM79 | ( | double | T_gas, |
| double | T_grain | ||
| ) |
Definition at line 4460 of file mole_reactions.cpp.
References DEBUG_ENTRY, and S.
Referenced by sticking_probability_H_func().
|
static |
Definition at line 82 of file mole_reactions.cpp.
|
static |
Definition at line 2828 of file mole_reactions.cpp.
 1.8.5
1.8.5